
Table des matières
RAPIDS: Accelerated data science on GPUs
This page highlights the RAPIDS(NVIDIA) accelerated solution on GPUs for all data science jobs and Big Data: data analysis, data visualization, data modelization, machine learning (random forest, PCA, gradient boosted trees, k-means, …). On Jean Zay, you simply need to call the module to use RAPIDS; for example:
module load rapids/21.08
The most suitable solution for a data study with RAPIDS on Jean Zay is to open a notebook via a dynamic allocation on a compute node using our JupyterHub instance.
On a compute node, there is no connection through internet. All library installations or data downloading should be done before the dynamic allocation on a compute node.
At the end of this page, you will find a link to the example notebooks of RAPIDS functionalities.
Comment: This documentation is limited to the use of RAPIDS on a single node. In a Big Data context, if you need to distribute your study on more than 8 GPUs in order to accelerate your study even more, or because the size of the DataFrame handled exceeds the memory of all the GPUs (more than 100 GB), the IDRIS support team can assist you in configuring a cluster on Jean Zay with the dimensions your usage.
GPU acceleration
The use of RAPIDS and GPUs becomes advantageous, compared to classic solutions on CPUs, when the data table being manipulated reaches multiple gigabytes of memory. In this case, the acceleration of most of the read/write tasks, data preprocessing and machine learning is significant.
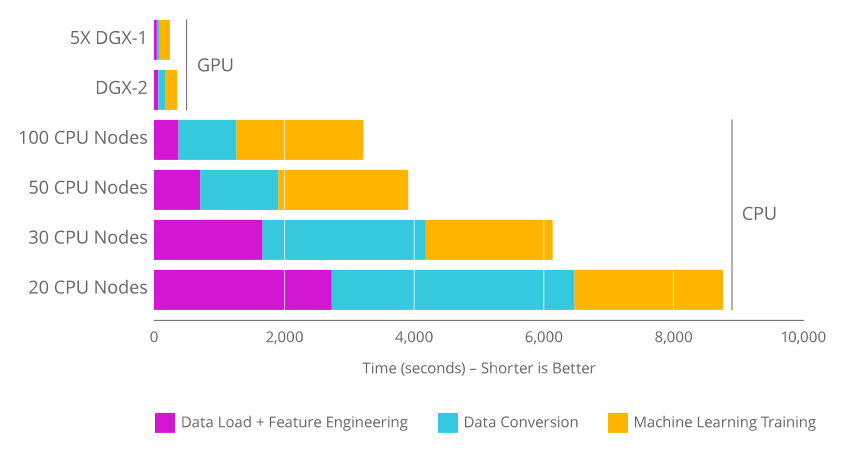
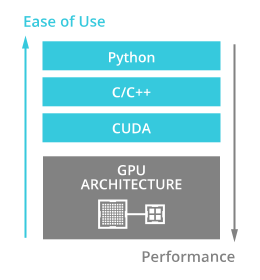
The RAPIDS APIs optimize the use of GPUs for data analysis jobs and machine learning. The Python code porting is easy to use: All the RAPIDS APIs reproduce the legacy API functionalities in CPU processing.
Platform and RAPIDS API
RAPIDS is based on the Apache Arrow project which manages the memory storage of data structures in columns.
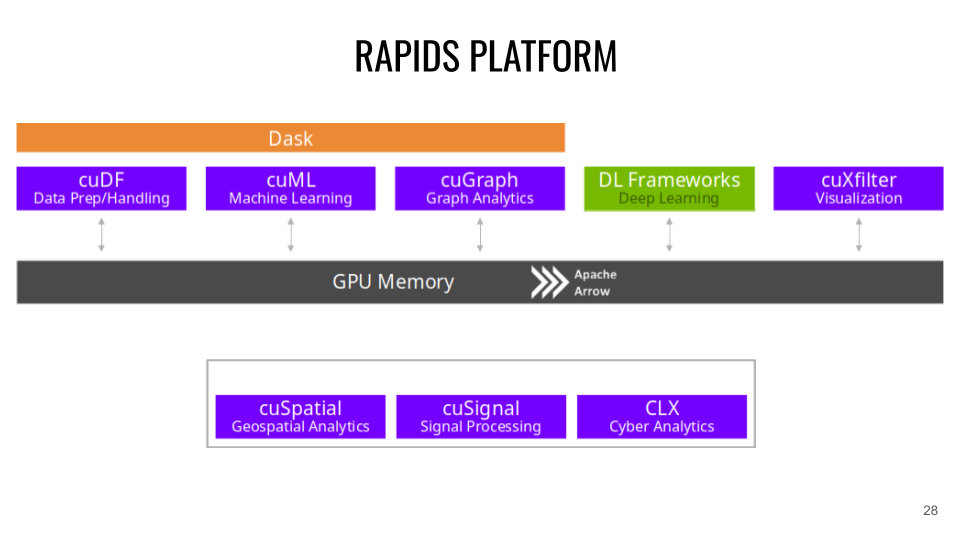
* CuDF (CPU communication: Pandas ). CuDF is a GPU DataFrame library for data loading, fusion operations, aggregation, filtering and other data handling. Example:
import cudf tips_df = cudf.read_csv('dataset.csv') tips_df['tip_percentage'] = tips_df['tip'] / tips_df['total_bill'] * 100 # display average tip by dining party size print(tips_df.groupby('size').tip_percentage.mean())
* CuML (CPU communication: scikit-learn ). CuML Machine Learning algorithm library on tabular data compatible with the other RAPIDS APIs and the Scikit-learn API. (XGBOOST can also be directly used on the RAPIDS APIs). Example:
import cudf from cuml.cluster import DBSCAN # Create and populate a GPU DataFrame gdf_float = cudf.DataFrame() gdf_float['0'] = [1.0, 2.0, 5.0] gdf_float['1'] = [4.0, 2.0, 1.0] gdf_float['2'] = [4.0, 2.0, 1.0] # Setup and fit clusters dbscan_float = DBSCAN(eps=1.0, min_samples=1) dbscan_float.fit(gdf_float) print(dbscan_float.labels_)
* CuGraph (CPU communication: NetworkX ). CuGraph is a Graph analysis library which uses a GPU Dataframe input. Example:
import cugraph # read data into a cuDF DataFrame using read_csv gdf = cudf.read_csv("graph_data.csv", names=["src", "dst"], dtype=["int32", "int32"]) # We now have data as edge pairs # create a Graph using the source (src) and destination (dst) vertex pairs G = cugraph.Graph() G.from_cudf_edgelist(gdf, source='src', destination='dst') # Let's now get the PageRank score of each vertex by calling cugraph.pagerank df_page = cugraph.pagerank(G) # Let's look at the PageRank Score (only do this on small graphs) for i in range(len(df_page)): print("vertex " + str(df_page['vertex'].iloc[i]) + " PageRank is " + str(df_page['pagerank'].iloc[i]))
* Cuxfilter (CPU communication: Bokeh / Datashader ). Cuxfilter is a data visualization library. It enables connecting a GPU-accelerated crossfiltering to a web depiction.
* CuSpatial (CPU communication: GeoPandas / SciPy.spatial ). CuSpatial is a spatial data processing library including point-in-polygons, spatial joins, geographical coordinate systems, primitive shapes, trajectory distances, and analyses.
* CuSignal (CPU communication: SciPy.signal ). CuSignal is a signal processing library.
* CLX (CPU communication: cyberPandas ). CLX is a library of processing and cybersecurity data analysis.
Multi-GPU acceleration with Dask-CUDA
The Dask-CUDA library enables using the API RAPIDS principals on multiple GPUs. This enables the acceleration of your code even more, and above alll to be sable to process the data tables of several tens or hundreds of gigabytes which would surpass the RAM memory of a single GPU.
After having configured a cluster, Dask functions as a workflow. Each called operation builds a computational graph. It is only when the computation is commanded that it is effectuated.
from dask.distributed import Client from dask_cuda import LocalCUDACluster cluster = LocalCUDACluster() client = Client(cluster)
The graph computation is started with the .compute() command. The result will be stored in the memory of a single GPU.
import dask_cudf ddf = dask_cudf.read_csv('x.csv') mean_age = ddf['age'].mean() mean_age.compute()
Typically, in a Dask workflow where several successive actions are applied to the data, it would be faster to persist the data in the cluster (to keep the data parts which are distributed and associated to each GPU in the memory of each GPU). For this, the .persist() command is used.
ddf = dask_cudf.read_csv('x.csv') ddf = ddf.persist() df['age'].mean().compute()
Example avec Cuml:
from dask.distributed import Client from dask_cuda import LocalCUDACluster import dask_cudf import cuml from cuml.dask.cluster import KMeans cluster = LocalCUDACluster() client = Client(cluster) ddf = dask_cudf.read_csv('x.csv').persist() dkm = KMeans(n_clusters=20) dkm.fit(ddf) cluster_centers = dkm.cluster_centers_ cluster_centers.columns = ddf.column labels_predicted = dkm.predict(ddf) class_idx = cluster_centers.nsmallest(1, 'class1').index[0] labels_predicted[labels_predicted==class_idx].compute()
Example notebooks on Jean Zay
You can find the example notebooks (cuml, xgboost, cugraph, cusignal, cuxfilter, clx) in the RAPIDS demo containers.
To copy the notebooks into your personal space, you need to execute the following command:
cp -r $DSDIR/examples_IA/Rapids $WORK
